Introduction
The first antibody libraries were generated using phage display technology in the 1990s since the conception of phage display technology in 19851, which allowed for the display of antibodies on the surface of bacteriophages2. Since then, various other display technologies have been developed, including yeast display (19973), ribosome display (19944, 19975) and mammalian display. These technologies have allowed for the generation of larger and more diverse antibody libraries, as well as the selection of specific antibodies with improved properties, providing a powerful tool for the discovery and engineering of antibodies.
Phage display
Phage display is a widely used in vitro selection method for antibodies displayed on the surface of bacteriophages binding to specific ligands. It allows for rapid screening of a large number of antibody fragments in a single experiment, making it a highly efficient method for antibody discovery.
Phage display is relatively simple to use and does not require specialized equipment or extensive cell culture conditions. It can be used to identify antibodies against a wide range of antigens, including proteins, peptides, and small molecules.
The technology also does not require a significant investment in equipment or reagents, making it relatively affordable. The stability of phages is another advantage and they can be stored for extended periods without losing their ability to display antibodies. Some of the limitations of phage display technology include the risk of phage contamination in downstream applications.
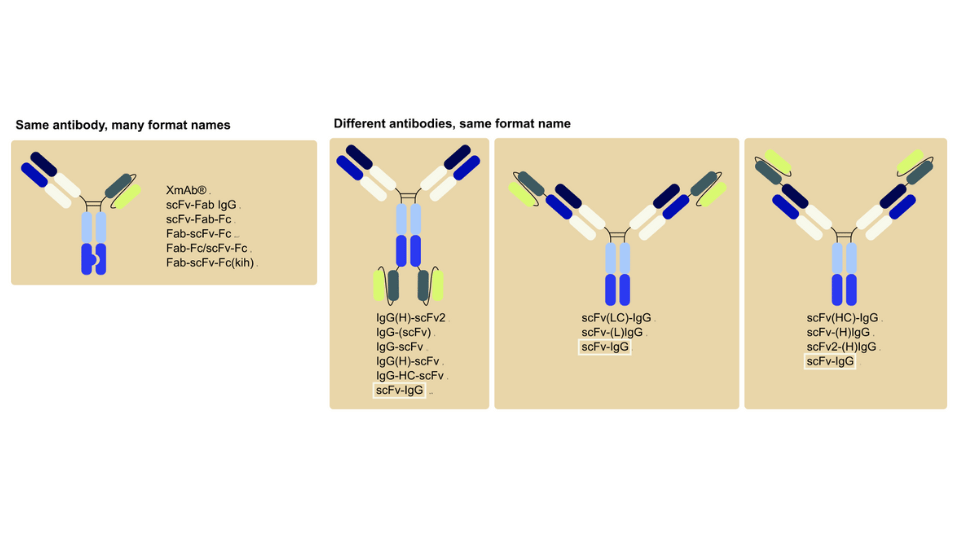
Especially scFv (single-chain variable fragments) and Fab (antigen-binding fragments) formats are used for display, with scFvs often providing a more robust format for expression7. Reformatting scFv and Fab formats into IgGs (providing longer half-life and triggers for effector functions) is also practical.

Also single-domain VHH antibodies are used for phage display. Just one recent example of a high-throughput sequencing-based strategy from Hanke et al. (2022) included combining phage display of VHH, NGS and multivariate enrichment analysis with functional testing to identify and isolate cross-neutralizing SARS-CoV-2 nanobodies8.
The fundamental importance of phage display technology was recognized in 2018 by awarding the Nobel Prize in Chemistry to George Smith and Sir Gregory Winter.
Mammalian display
Mammalian surface display is a newer technology for generating antibody libraries and selecting specific antibodies. In this method, antibodies are expressed as fusion proteins in mammalian cells and can be selected using various techniques, including FACS, magnetic bead-based selection, and biopanning.
One of the main advantages of mammalian display is expression antibodies in their native form. Full-length Ig molecules can be displayed on the cell surface, which makes the technology also suitable also for production.
The limited size of mammalian display libraries is one of the largest drawbacks of the method, but even relatively large antibody libraries have been displayed on mammalian cells. For example, Parthiban et al. (2019)9 designed an IgG library of 107 CDR3 mutated clones for affinity maturation in order to identify antibody clones with better PD-1-blocking properties.
Yeast display
The yeast surface display technique was originally developed by Boder and Wittrup in 19973. In yeast display, antibodies are typically expressed in yeast cells (S. cerevisiae) as scFvs and Fabs, but even full-length Igs (IgG1 library in Rouha et al.10 and IgGs in Rhiel et al.11). The two proteins, Aga1 and Aga2, are typically employed as anchors between the antibody fragment and yeast cell12.

Antibody fragments can be selected using various techniques, normally fluorescence-activated cell sorting (FACS), magnetic bead-based selection, and NGS-based approaches. Affinity maturation can be performed through biopanning.
Yeast display offers some advantages, including its suitability for in vivo applications and its ability to express antibodies in their native form, while yeast display libraries are typically smaller in size, compared to phage display, for example.
Bacterial display
In bacterial display, antibody fragments can be displayed on the surface of bacterial cells, commonly with lipoproteins, such as the Lpp-OmpA. Bacteria such as E. coli is commonly used for peptide display13 and, for example, for displaying antigens in epitope mapping14. In addition to this, E. coli 15, 16 and also gram positive bacteria belonging to the Staphylococcus genus (xylosus17, aureus18, carnosus19) have been used for antibody display. Displayed formats include full-length IgGs, fragments and VHH.
Post-translational modifications unlike those in human cells as well as lower transformation efficiencies (for gram positive bacteria) are some of the drawbacks of the display method. However, large library sizes and fast growth rates make bacterial cells a valid option for in vitro display.
Ribosome display
Ribosome display is a cell-free display system that does not require the use of living cells for antibody expression and selection. Instead, it relies on cell-free protein synthesis reactions to produce and display antibody fragments. Display methods for large, diverse libraries of multi-chain fragments for display and selection have been developed for ribosome display. In an method by Stafford et al. (2014), Fab fragments could even be expressed as full-length IgGs20.

Ribosome display is not limited by host cell transformation efficiency which allows large libraries of up to 1012 and 1015 antibody variants. It is also fast as no cell culture is required. However, lower levels of accessible functional ribosomes are one limitation of the technology21.
Summary
The cell and cell-free display technologies presented above have enabled rapid discovery of novel antibody fragments entirely in vitro.
Phage display remains the most widely used in vitro display technology due to its ease of use and versatility. Mammalian display offers several advantages, such as post-translational modifications, that are lacking in other display platforms. Ribosome display allows for cell-free synthesis of antibodies, while yeast display allows for native expression of antibodies. Bacterial display, although less commonly used, offers advantages such as low cost and ease of expression.
In vitro display technologies have led to the development of several clinically approved therapeutic antibodies, and their use in research and therapeutic applications is expected to increase. Improved library designs, increased library sizes, faster and more effective screening methods, as well as the development of technologies for the selection of antibodies against membrane proteins and other difficult targets will surely remain areas of interest in future research.